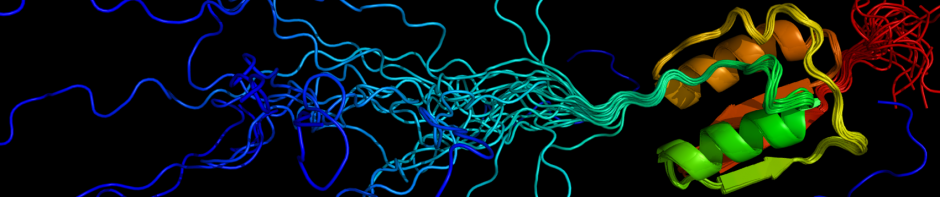
Our main interest is to the study the structural and functional properties of intrinsically disordered proteins and their involvement in various diseases. We develop various computational tools to aid research in this field, including prediction tools (e.g. IUPred and IUPred2A, ANCHOR) and databases (e.g. DIBS, PhaSePro). As many disordered proteins function through short linear motifs, we are also interested in how such functional sites can be identified from the amino acid sequence. We also study specific linear motif systems using a combination of computational and experimental methods. In particular, we are exploring the interaction partners of the dynein light chain LC8, and specific E3 ligases, such as SIAH.
We are coordinating the IDP2Biomed twinning rant. Our group is also involved in EU funded projects IDPro and IDPFun2. We are coordinating the Twinning grant starting in October 1st, 2024 with partners Prof. Michele Vendruscolo (University of Cambridge), Prof. Markus Zweckstetter (German Center for Neurodegenerative Diseases), Prof. Silvio Tossato (University of Padova).
Selected publications
- AIUPred: combining energy estimation with deep learning for the enhanced prediction of protein disorder
Gábor Erdős, Zsuzsanna Dosztányi
Nucleic Acids Research, Volume 52, Issue W1, 5 July 2024, Pages W176–W181, https://doi.org/10.1093/nar/gkae385 - Tutorial: a guide for the selection of fast and accurate computational tools for the prediction of intrinsic disorder in proteins
L. Kurgan, G. Hu, K. Wang, S. Ghadermarzi, B. Zhao, N. Malhis, G. Erdős, J. Gsponer, V. N. Uversky, Zs. Dosztányi
Nature Protocols, 2023, 10.1038/s41596-023-00876-x - DisCanVis: Visualizing integrated structural and functional annotations to better understand the effect of cancer mutations located within disordered proteins
Norbert Deutsch, Mátyás Pajkos, Gábor Erdős, Zsuzsanna Dosztányi
Protein Science, 2023 Jan;32(1):e4522. doi: 10.1002/pro.4522. - The interaction between LC8 and LCA5 reveals a novel oligomerization function of LC8 in the ciliary-centrosome system
Tamás Szaniszló, Máté Fülöp, Mátyás Pajkos, Gábor Erdős, Réka Ágnes Kovács, Henrietta Vadászi, József Kardos & Zsuzsanna Dosztányi
Scientific Reports volume 12, Article number: 15623 (2022) - Deep Learning for Intrinsically Disordered Proteins: From Improved Predictions to Deciphering Conformational Ensembles
Gábor Erdős, Zsuzsanna Dosztányi
Current Opinion in Structural Biology, Volume 89, December 2024, 102950
Services
Funding
 |  |  |  |  |